News
Artist's rendition of transcription and translation in cells. Lin and Amir propose a new model for gene expression, where competition for polymerases/ribosomes are introduced into the transcription and translation processes. This competition accounts for the exponential growth of protein levels in growing cells and leads to protein level homeostasis. (Image credit: Equinox Graphics Ltd.)
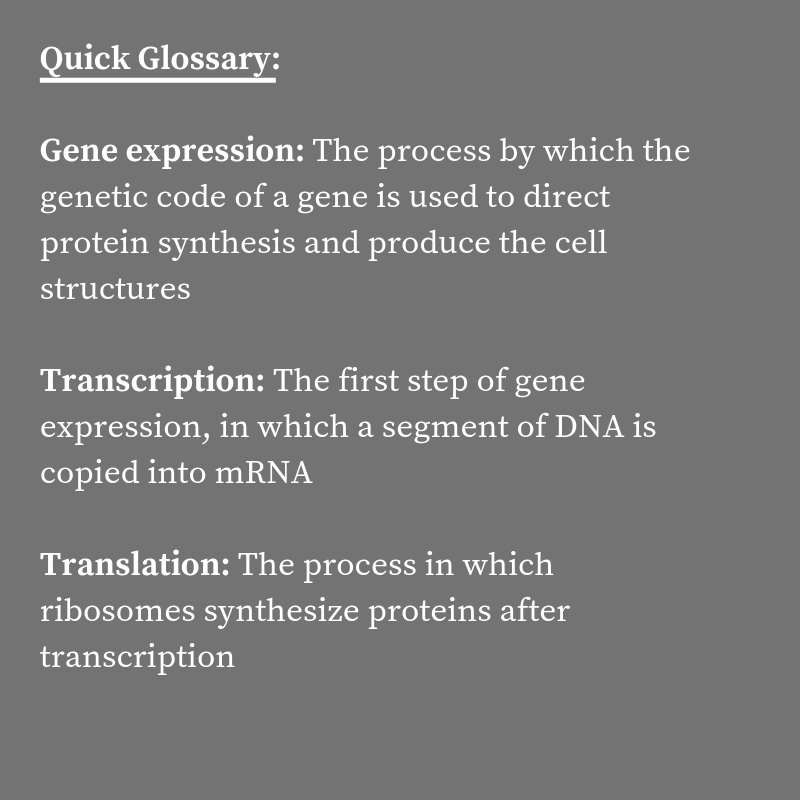
Researchers at the Harvard John A. Paulson School of Engineering and Applied Sciences (SEAS) have developed a mathematical model that sheds new light on the fundamental processes of gene expression in cells.
While the primary players in gene expression — DNA, mRNA, ribosomes, and polymerases — have been known since the 1950s, how they interact during the cell cycle is still unclear.
In previous models, researchers thought that the rate of transcription (making a mRNA copy of DNA) and translation (turning mRNA into a sequence of amino acids) was limited by the amount of DNA and mRNA available in the cell.
But the new model, combined with existing experimental data, actually suggests that ribosomes and polymerases control the levels of proteins and mRNA in the cell.
“Our research shows that widely used models for gene expression and various other cellular processes have to be revised in an interesting way,” said Ariel Amir, Assistant Professor in Applied Mathematics at SEAS and senior author of the paper.
The research was published recently in Nature Communications.

To understand how the cell cycle works, think of the cell like a popular restaurant that needs to constantly add new tables. The question is: How many tables can be added before the kitchen can’t keep up?
All of the restaurant’s menu items come from a family cookbook, the DNA. Inside that cookbook are a number of recipes, the mRNA. Those recipes get copied by a Xerox machine, polymerases, and cooked by chefs, ribosomes.
In the previous model of gene expression, the growth of the restaurant is limited by the number of cookbooks, which limits the number of recipes floating around at any point in time.
The new model, however, says the growth of the restaurant is often limited by the number of Xerox machines copying recipes and the number of chefs in the kitchen. It is the ribosomes and polymerases that limit gene expression.
“Since gene expression is fundamental in biology, the new model could have implications on our understanding of other cellular processes, including cell cycle regulation, cell shape regulation and more,” said Jie Lin, the George Carrier Postdoctoral Fellow in Applied Mathematics at SEAS and first author of the paper.
This research was supported by the Alfred P. Sloan Foundation, the Milton Fund, the Volkswagen Foundation, the Harvard Deans Competitive Fund for Promising Scholarship, and the National Science Foundation through the Harvard Materials Research Science and Engineering Center.
Topics: Applied Mathematics
Cutting-edge science delivered direct to your inbox.
Join the Harvard SEAS mailing list.
Scientist Profiles
Ariel Amir
Associate in Applied Mathematics
Press Contact
Leah Burrows | 617-496-1351 | lburrows@seas.harvard.edu