News
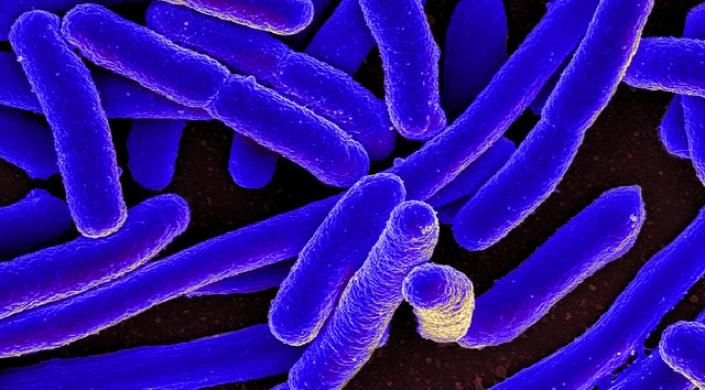
Colorized scanning electron micrograph of Escherichia coli, grown in culture and adhered to a cover slip (Image courtesy of the NIH)
Bacteria come in all shapes and sizes — some are straight as a rod, others twist like a corkscrew. Shape plays an important role in how bacteria infiltrate and attack cells in the body.
The helical shape of Helicobacter pylori, a species of bacteria which can cause ulcers, may help it penetrate tissues.
Bacteria have an extraordinary ability to maintain and recover their morphology even after being twisted out of shape. Researchers know that shape is determined by the cell wall, yet little is known about how bacteria monitor and control it. Since the cell wall is the target of most antibiotics, understanding how bacteria grow their cell walls may provide insight into more effective medicines.
Now, a team of researchers led by the Harvard John A. Paulson School of Engineering and Applied Sciences (SEAS) has found that Escherichia coli (E. coli) may use mechanical cues to keep their shape.
The research is published in Nature Microbiology.
“This research may reveal some basic principles of bacteria growth,” said Felix Wong, a graduate student at SEAS and co-first author of the paper. “We showed that the coupling of cell wall growth to mechanical strain is quantitatively consistent with how bacteria recovered their shape after being deformed in experiments.”

Stressed-out E.coli recovers its straight, rod-like shape over time. (Images courtesy of Lars D. Renner at the Leibniz Institute of Polymer Research and the Max Bergmann Center of Biomaterials, Dresden, Germany.)
Wong and senior author Ariel Amir, assistant professor in Applied Mathematics, began by modeling the mechanics of the E. coli cell wall under constraints that forced the bacteria to grow into the shape of a donut.
In previous research, Amir observed that under similar bending forces bacteria become plastically deformed, meaning when the bending force was removed, E. coli cells snapped back to a straighter, but still bent, shape. This suggested that cell wall growth could sense the applied bending force. Amir also found that the cells straightened upon further growth, an observation which was left unresolved in that paper.
In the latest research, the team explored whether coupling wall growth to mechanical strain — how the bacterium was compressed or stretched — could explain the snapback and predict how fast the bacteria would straighten when released.
Wong and Amir answered this question with a theoretical model which quantitatively predicted both how the bacteria would grow to recover its straight shape and how long it would take.
Then, together with the experimental groups of Drs. Lars Renner and Sven van Teeffelen, from the Leibniz Institute of Polymer Research and the Max Bergmann Center of Biomaterials in Germany and the Institut Pasteur in France, respectively, ran the experiment with E. coli.
The models and experiments were consistent with each other. A mechanical strain-dependent cell wall growth rate predicted a straightening rate consistent with what was found experimentally.
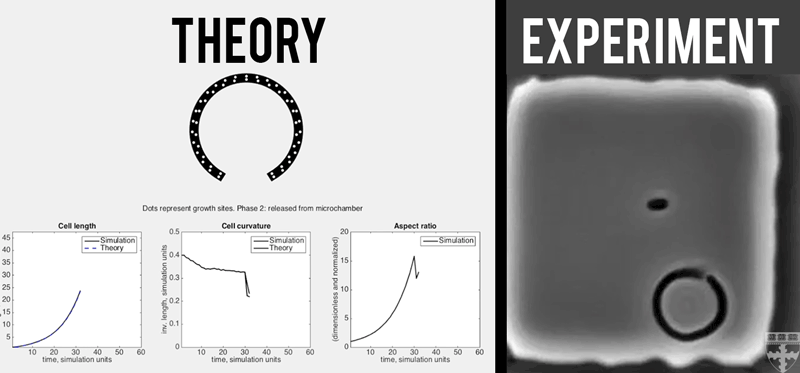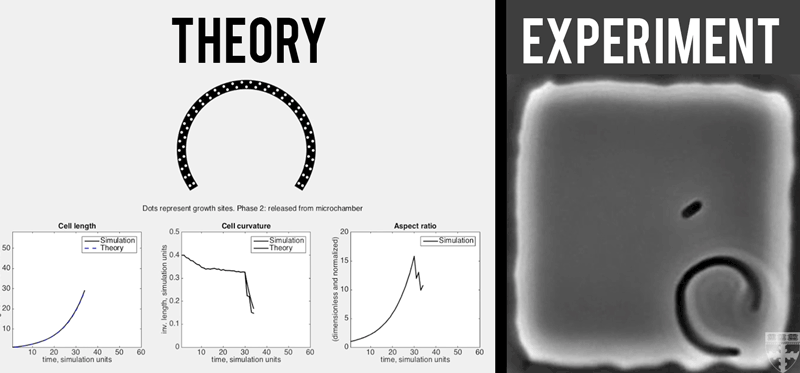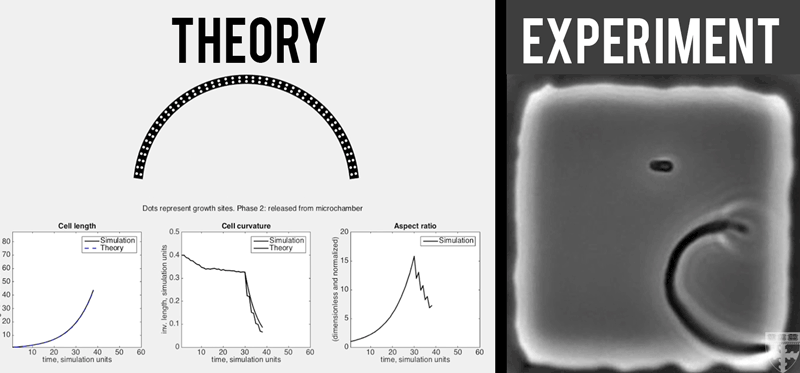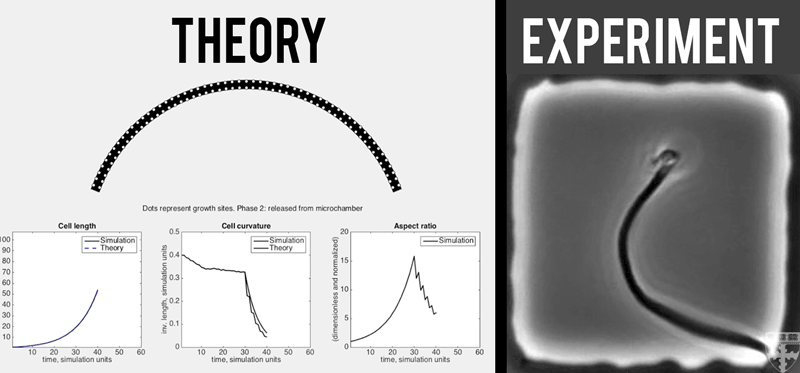
A theoretical model (left) quantitatively predicts how E. coli (right) grown in confined, micro-chambers recover their straight, rod-like morphologies overtime. (Video courtesy of Harvard SEAS)
“We think our proposed idea for bacteria is reminiscent of plant growth,” said Wong. “It’s been well established in the plant field that mechanical cues can bias plant growth. Our research shows that the same may well be true for bacteria. However, if mechanical strains were indeed an important sensory cue for bacteria, then there has to be a molecular mechanism that senses mechanical strain.”
Next, the researchers hope to identify and understand those molecular mechanisms, which could provide new targets for antibiotics in the future.
The paper was co-authored by Gizem Özbaykal, Jayson Paulose and Douglas B. Weibel. The research was supported in part by the National Science Foundation, the Volkswagen Foundation and the Alfred P. Sloan Foundation.
Cutting-edge science delivered direct to your inbox.
Join the Harvard SEAS mailing list.
Scientist Profiles
Ariel Amir
Associate in Applied Mathematics
Press Contact
Leah Burrows | 617-496-1351 | lburrows@seas.harvard.edu